A college reached out yesterday about a performance issue they were hitting when working with the Microsoft Building Footprints dataset we host on the Planetary Computer. They wanted to get the building footprints for a small section of Turkey, but noticed that the performance was relatively slow and it seemed like a lot of data was being read.
This post details how we debugged what was going on, and the steps we took to fix it.
The problem
First, my college sent a minimal, complete, and verifiable example of the problem. This let me very easily reproduce it. From his report, the first thing I suspected was an issue with the spatial partitioning. The files were supposed to be partitioned by quadkey, so that all the building footprints in a single area are in the same partition. Then spatial queries will be very fast: you only need to load a small subset of the data.
When I benchmarked things, it took about:
- 16 seconds to read the metadata with
dask_geopandas.read_parquet - 60 seconds to read the data and clip it to the area of interest
Looking at the spatial partitions of the data showed that it was clearly not spatially partitioned:
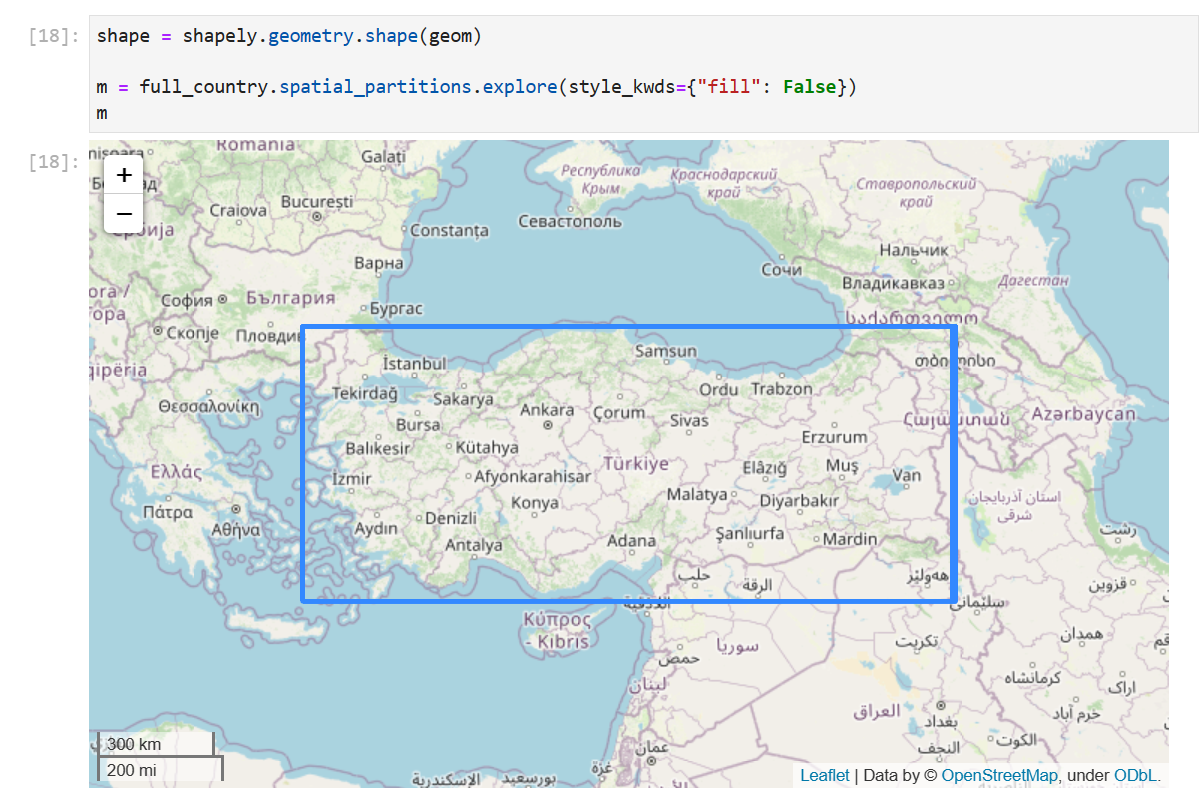
It’s clearer zoomed in, but the box is a bit fuzzy because it’s actually a bunch of boxes with very slightly different extents.
Turns out we dropped a few of the newer ms-buildings STAC items, which were spatially partitioned, during our last release. Oops. (Don’t worry, we’re working on a better system for this.)
Once I got those items re-ingested, things did look better.
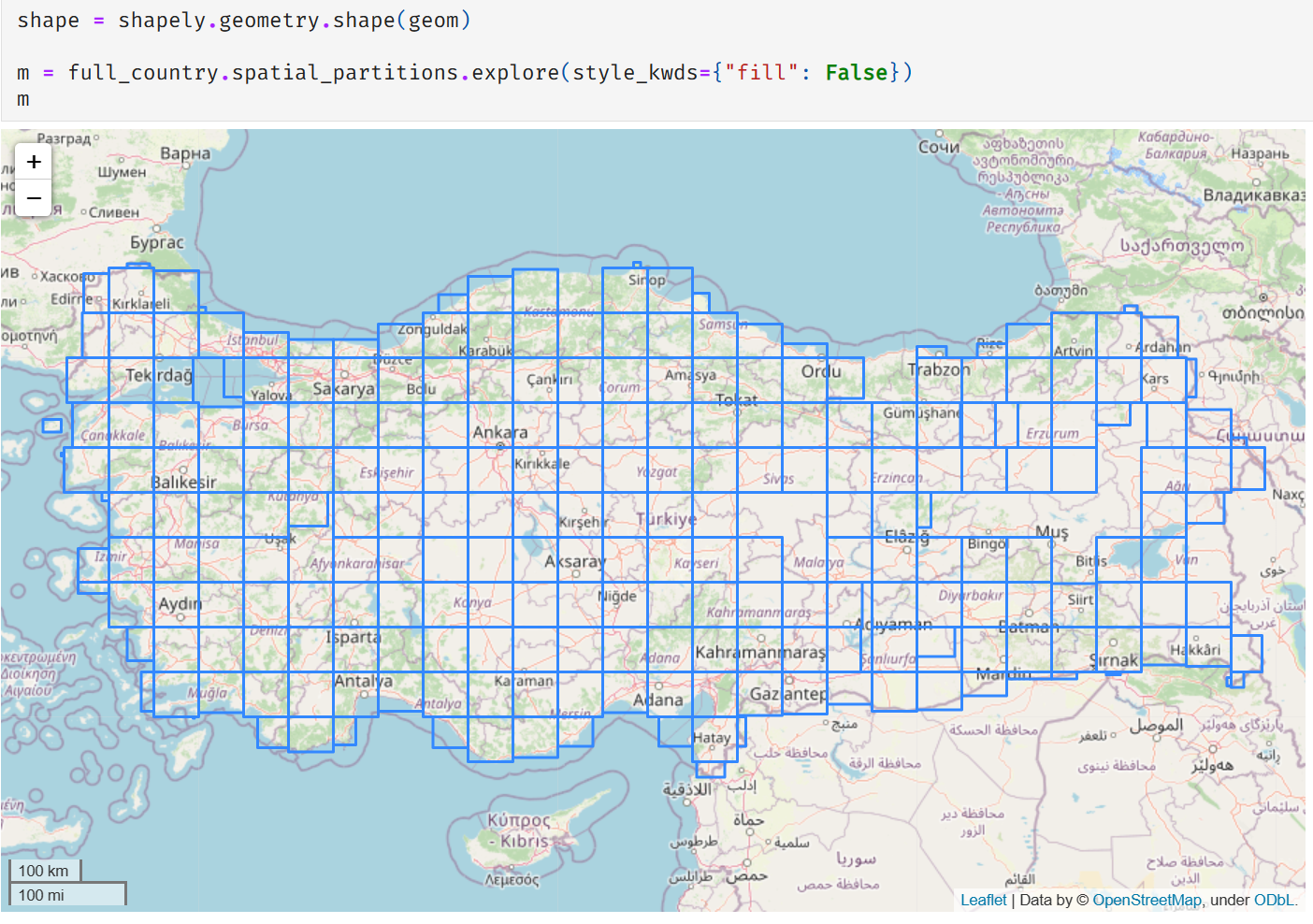
It wasn’t all good news, though. Our timings went to
- 56 seconds to read the metadata with
dask_geopandas.read_parquet(ouch) - 0.5 seconds to read the data and clip it to the area of interest (yay!)
The speedup from 60 seconds to 0.5 seconds is exactly why we want to spatially partition the data. When you’re querying for a small area of interest, the spatially partitioned data means you can ignore most of the data and speed things up a lot. But what’s going on with the slowdown for the first stage (reading metadata)?
The spatially partitioned dataset also had many more partitions in the Parquet
dataset, i.e. many more individual files in Blob Storage (a few hundred instead of 5-6).
At the moment, dask-geopandas needs to open each individual file to read its spatial bounds. That was fine
when we only had a few files, but when you have a few hundred the small amount of time it takes to read
each file adds up. In this case, it added up to about 56 seconds of waiting just to read the metadata.
Speeding up the metadata reading
To speed up the metadata reading, we use the tried-and-true method of parallelizing it
with Dask (yes, we’re using Dask to speed up Dask). Instead of doing a dask_geopandas.read_parquet
on the client (which in turn executes some pyarrow.parquet stuff to read the fragments and get
the metadata from each file) in serial, we’ll run a bunch of dask_geopandas.read_parquet
calls on the cluster in parallel (I’m just using a LocalCluster in this example).
The snippet below re-uses dask_geopandas.read_parquet, but applies it in
parallel using client.map. We’ll make a bunch of Dask DataFrames on the
cluster (one per file) and then we use client.gather to bring back the Dask
DataFrames (just the metadata, not the data!) to the client and concat them
together into one big Dask DataFrame.
There’s a small bug in dask-geopandas around serializing the spatial
partitions on a Dask GeoDataFrame. Once my fix is merged then this will
be a bit cleaner: everything to do with spatial_partitions can be deleted
and you’re just left with reading the metadata on the cluster, bringing it
back to the client, and concatenating at the end.
import distributed
import dask_geopandas
import dask.dataframe as dd
import fsspec
def read_parquet(paths, storage_options):
client = distributed.get_client()
# Read each partition's metadata on the cluster
df_futures = client.map(
dask_geopandas.read_parquet, paths, storage_options=storage_options
)
# workaround https://github.com/geopandas/dask-geopandas/issues/237
def get_spatial_partitions(x):
return x.spatial_partitions
spatial_partitions_futures = client.map(get_spatial_partitions, df_futures)
# Pull back locally. This takes the most time, waiting for computation
dfs, spatial_partitions = client.gather([df_futures, spatial_partitions_futures])
for df, sp in zip(dfs, spatial_partitions):
df.spatial_partitions = None
full_country = dd.concat(dfs)
full_country.spatial_partitions = pd.concat(spatial_partitions, ignore_index=True)
return full_country
We get the paths with something like
>>> fs, token, [root] = fsspec.get_fs_token_paths(asset.href, storage_options=asset.extra_fields["table:storage_options"])
# Get the raw paths (fast enough to do this locally. Could be done on the cluster too)
>>> paths = [f"az://{p}" for p in fs.ls(root)]
Note that there’s an open issue on Dask to do this kind of thing by default.
Overall, we brought the metadata read time down the 30 seconds (which would be faster with more workers). Still not great, but an improvement. At some point we’ll need to embrace a broader solution to this metadata access issue using something like Apache Iceberg.
See this example notebook for the full thing.